Stop Claude from Hallucinating Synthea Modules
LLMs generate valid Synthea JSON but hallucinate the medical codes. Here's a Skill to grounds every SNOMED and LOINC lookup.
mock.health · 14 min read · 2026-04-08
Synthea has 85 disease modules. Each one is a JSON state machine that generates encounters, conditions, labs, medications, and procedures for a specific disease. If you need a condition Synthea doesn't cover — celiac disease, migraine, GERD, whatever — you author a new module.
The module format is learnable. The hard part is the medical codes.
Every module embeds SNOMED codes for conditions, LOINC codes for labs, and RxNorm codes for medications. Ask an LLM to write a celiac disease module and it'll generate SNOMED code 396331005. That's correct. Ask it for a duodenal biopsy and it might generate 12866006. Looks right. Validates as a real SNOMED code. It's actually pneumococcal vaccination.
You can't tell a valid code from a hallucinated one by looking at it. The only way to know is to check it against a terminology server. This post teaches that workflow, and we published a Claude Code skill that automates it.
Why You'd Want Custom Modules
We generated 10,000 patients from a fresh Synthea clone and compared against CDC benchmarks. The results explain why the built-in modules aren't enough:
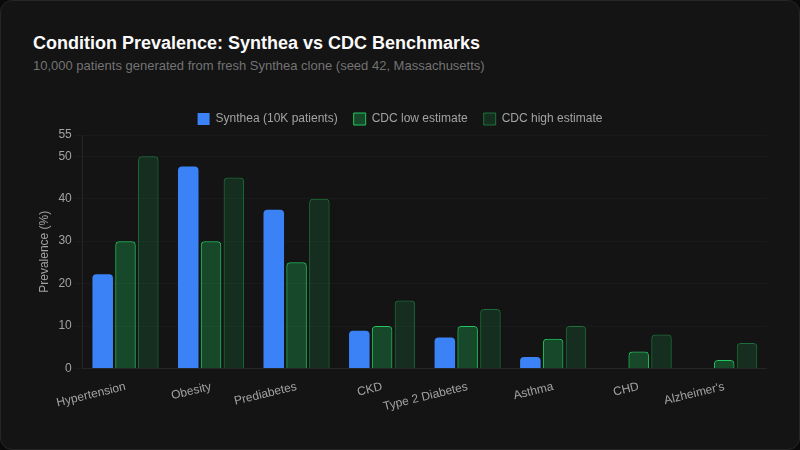
Coronary heart disease: 0%. Alzheimer's: 0%. Entire disease categories missing from 10,000 patients. We wrote about the architecture behind this in how we validate synthetic data — the short version is that each module runs independently, so conditions don't interact the way real diseases do.
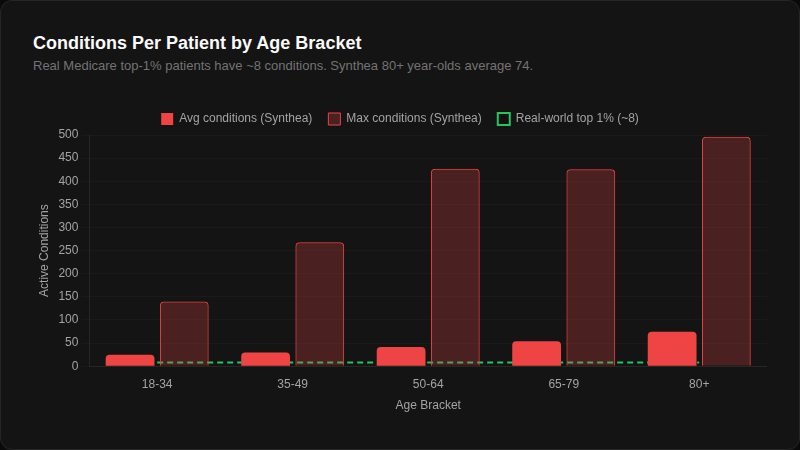
An average 80+ year-old Synthea patient has 74 active conditions. The top 1% of real Medicare patients have about 8. Most of Synthea's "conditions" are social determinants and administrative codes — Medication review due (situation) appears in 100% of patients. The green dashed line showing the real-world benchmark is barely visible because the scale is that far off.
If your use case needs a disease that Synthea's 85 modules don't cover, or needs more realistic disease modeling than the built-in coin-flip architecture provides, you're authoring a module.
The Module Format in 60 Seconds
A Synthea module is a JSON file with a name, a states object, and a gmf_version. Each state has a type and a transition to the next state. Here's the smallest useful module — a condition that gets diagnosed at an encounter:
{
"name": "Example Condition",
"states": {
"Initial": {
"type": "Initial",
"distributed_transition": [
{ "distribution": 0.01, "transition": "Onset" },
{ "distribution": 0.99, "transition": "Terminal" }
]
},
"Onset": {
"type": "ConditionOnset",
"codes": [{ "system": "SNOMED-CT", "code": "??????", "display": "??????" }],
"direct_transition": "Terminal"
},
"Terminal": { "type": "Terminal" }
},
"gmf_version": 2
}
The ?????? is the problem. What SNOMED code goes there? You need to look it up.
The state types you'll actually use: Initial, Terminal, Encounter/EncounterEnd (start and end clinical visits), ConditionOnset/ConditionEnd (diagnose and resolve), MedicationOrder/MedicationEnd (prescribe and stop), Observation (labs and vitals), Procedure (medical procedures), Guard (wait for a condition to be true), Delay (time passage), and SetAttribute (store patient variables).
Transitions come in four flavors: direct_transition (always go here), conditional_transition (if-then branching), distributed_transition (probability-weighted), and complex_transition (conditions with probability distributions). Real modules use all four.
The Code Problem
Every clinical state in a Synthea module needs codes from standard vocabularies:
| System | What it codes | Example |
|---|---|---|
| SNOMED-CT | Conditions, procedures, findings | 396331005 = Celiac disease |
| LOINC | Lab results, vital signs | 31017-7 = tTG IgA antibody |
| RxNorm | Medications | 310325 = Ferrous sulfate 325mg |
LLMs pattern-match these codes from training data. They don't look them up. For common conditions the codes are usually right — there's enough training signal. For anything less common, the LLM generates a plausible-looking number that might not exist in the code system at all.
The fix: tx.fhir.org, a free public FHIR terminology server maintained by HL7. No account needed. One curl call validates any code:
# Is SNOMED 396331005 a real code?
curl -s "https://tx.fhir.org/r4/CodeSystem/\$validate-code?\
system=http://snomed.info/sct&code=396331005" \
| jq '.parameter[] | select(.name=="result" or .name=="display")'
{ "name": "result", "valueBoolean": true }
{ "name": "display", "valueString": "Coeliac disease" }
And when you need to find the right code:
# Search SNOMED for "celiac disease"
curl -s "https://tx.fhir.org/r4/ValueSet/\$expand?\
url=http://snomed.info/sct?fhir_vs&filter=celiac+disease&count=5" \
| jq '.expansion.contains[] | {code, display}'
{ "code": "396331005", "display": "Coeliac disease" }
{ "code": "473213008", "display": "Celiac disease annual review" }
{ "code": "703970007", "display": "Celiac disease monitoring" }
This is what separates a working module from one that generates corrupt FHIR.
The Skill
We published a Claude Code skill that automates this workflow. It knows the module JSON schema, the code systems, and the validation endpoints. Install it and point it at a condition:
# Install the skill
claude install github:mock-health/samples/synthea-module-skill
# Use it
claude "/synthea create a celiac disease module"
The skill follows a six-step workflow:
- Check existing modules — searches
src/main/resources/modules/to avoid duplicating what Synthea already has - Research the condition — prevalence rates, diagnostic criteria, treatment pathway
- Look up every code — validates each SNOMED, LOINC, and RxNorm code against tx.fhir.org before writing it into the module
- Generate the module JSON — following Synthea's exact schema with only validated codes
- Build and test — runs
./gradlew build -x testfor structural validation, then./run_synthea -m <name> -p 1to test generation - Inspect output — checks the generated FHIR bundle for expected resource types
The skill's SKILL.md contains the full module schema reference, code system mappings, grounding rules, and common pitfalls. It's the reference documentation the Synthea wiki doesn't have.
Working Example: Celiac Disease
Here's the module we built using this workflow. Every code was validated against tx.fhir.org before it went into the JSON.
Code inventory (all validated):
| Concept | System | Code | Display |
|---|---|---|---|
| Celiac disease | SNOMED-CT | 396331005 |
Coeliac disease |
| Encounter for problem | SNOMED-CT | 185347001 |
Encounter for problem |
| EGD | SNOMED-CT | 76009000 |
Esophagogastroduodenoscopy |
| Duodenal biopsy | SNOMED-CT | 235261009 |
Biopsy of duodenum |
| Gluten free diet | SNOMED-CT | 160671006 |
Gluten free diet |
| Diet education | SNOMED-CT | 11816003 |
Diet education |
| Iron deficiency anemia | SNOMED-CT | 87522002 |
Iron deficiency anemia |
| Follow-up encounter | SNOMED-CT | 390906007 |
Follow-up encounter |
| tTG IgA antibody | LOINC | 31017-7 |
Tissue transglutaminase IgA Ab [Units/volume] in Serum |
| Ferritin | LOINC | 2276-4 |
Ferritin [Mass/volume] in Serum or Plasma |
| Ferrous sulfate | RxNorm | 310325 |
ferrous sulfate 325 MG (iron 65 MG) Oral Tablet |
The state machine:
Initial → Age_Guard (wait until age 2)
→ Prevalence_Check (monthly, age-stratified probability)
→ Symptom_Onset → Diagnostic_Encounter
→ tTG IgA test → Referral_Delay (2-6 weeks)
→ Endoscopy_Encounter → EGD + Duodenal biopsy
→ Celiac_Diagnosis → Prescribe gluten-free diet
→ 50% chance: Iron deficiency → Prescribe ferrous sulfate
→ Annual monitoring loop (tTG IgA + ferritin)
The module uses complex_transition for age-stratified onset — childhood (higher rate) and ages 30-50 (second peak) get different probabilities. Newly diagnosed patients have a 50% chance of iron deficiency anemia (realistic for celiac). The monitoring loop runs annually with repeat serology and ferritin.
The full module JSON is about 200 lines. Drop it into synthea/src/main/resources/modules/celiac_disease.json and run:
cd synthea
./gradlew build -x test # validate structure
./run_synthea -m celiac_disease -p 10 -s 42 # generate 10 patients
jq -r '.entry[].resource.resourceType' output/fhir/*.json | sort | uniq -c | sort -rn
You'll get Encounters, Conditions (celiac disease, iron deficiency anemia), Observations (tTG IgA, ferritin), Procedures (EGD, duodenal biopsy), MedicationRequests (ferrous sulfate), and CarePlans (gluten-free diet).
For Deeper Work
/fhir Claude Code skill — for FHIR development beyond Synthea modules: R4/R5, Implementation Guides, FSH authoring, SMART on FHIR, validation.
Inferno — ONC's FHIR server compliance test suite. If you're testing whether your server conforms to US Core, this is the tool.
Synthea Module Builder — a GUI for visual module authoring. Good for understanding the state machine visually, but doesn't help with code grounding.
tx.fhir.org — the public FHIR terminology server. Free, no account. The skill uses it automatically, but you can query it directly for any code system.
The module format is learnable. The vocabulary problem is solvable. Ground your codes, validate your output, and don't trust any medical code an LLM generates from memory — including ours. We built mock.health because we hit the limits of module-level fixes. Population-level realism — comorbidity interactions, realistic prevalence, medication variety — requires a different architecture entirely. Free tier, API key in 60 seconds →
Related posts
- SMART on FHIR Tutorial: Build an App in 30 Minutes — Build a standalone SMART on FHIR app in a single HTML file with zero build tools. You can do it.
- Testing FHIR Integrations Without a Hospital — You can't get hospital access without a working integration. You can't build a working integration without hospital data. Here's how to break the catch-22.
- FHIR, USCDI, and US Core: What They Are, How They Fit — FHIR says how to send data. USCDI says what data. US Core says exactly how to format it. Here's how the three standards fit together.